|
Using standard sequencing techniques, only the most dominant DNA sequence can be identified, which means that in samples where more than one bacterial species is present (such as stool) results are uninterpretable. To reduce this risk, sequencing must be carried out to distinguish between a genuine pathogen and contaminants (often waterborne bacteria highly unlikely to cause disease). At a high number of thermal cycles, this low-level background contaminant DNA will be amplified and give a false-positive result. All bacterial DNA present in a sample is amplified, including that which is unavoidably present in reagents, meaning low-level environmental contamination is impossible to eliminate entirely. However, the breadth of broad-range 16S rDNA PCR renders it vulnerable to contamination. Broad-range 16S rDNA PCR enabled the identification of Bartonella henselae and Tropheryma whippelii as the pathogens underlying catscratch disease and Whipple’s disease, respectively. 8 16Īdditionally, broad-range 16S rDNA PCR may identify previously uncharacterised bacteria. 7 14 15 Broad-range 16S rDNA PCR, followed by sequencing, has allowed the differentiation and identification of both species, aiding research into species-specific pathogenicity. It is also possible to make distinctions between species: Ureaplasma spp consist of two bacterial strains, Ureaplasma parvum and Ureaplasma urealyticum, indistinguishable by culture, and each suspected to cause different pathology in neonates. 8 9 Examples include the identification of Helicobacter sp as the underlying cause of osteomyelitis or Neisseria meningitidis as the unexpected cause of septic arthritis by broad-range 16S rDNA PCR, after negative results were produced from other microbial diagnostic techniques. 8 11 The bacteria identified are often unusual, rare, difficult to culture, or bacteria for which a specific PCR is not available. It is also clinically useful when other techniques give negative results, for example, in culture-negative endocarditis, septic arthritis, meningitis or long-line infections. 5īroad-range 16S rDNA PCR can detect both viable and non-viable bacteria, similar to qPCR. Monitoring of the CT value can be used to assess therapeutic efficacy, for example, in tuberculosis. Strong positive results will be in the high teens or early 20s, whereas a sample with a CT value of 38–40 will be considered only borderline positive. The number of cycles taken to reach this threshold is known as the cycle threshold value (CT value) and is proportional to the initial quantity of DNA present in the sample: that is, the lower the CT value, the higher the initial quantity of DNA ( figure 1A). 4 The fluorescence is measured during the assay, and when it reaches a prespecified level the assay is considered positive. 3 In qPCR, the use of fluorescent probes enables bacterial load to be detected and quantified in real time, hence reducing time to diagnosis and correct treatment initiation. 2 Furthermore, some fastidious organisms may commonly escape detection in routine cultures, for example, Kingella kingae. In the future, analysis of individualised microbial communities using broad-range 16S rDNA PCR may be a key component of personalised medicine.īacterial culture takes at least 24–48 hours for determina tion of a positive result, or longer for slow-growing organisms such as Mycobacterium tuberculosis. 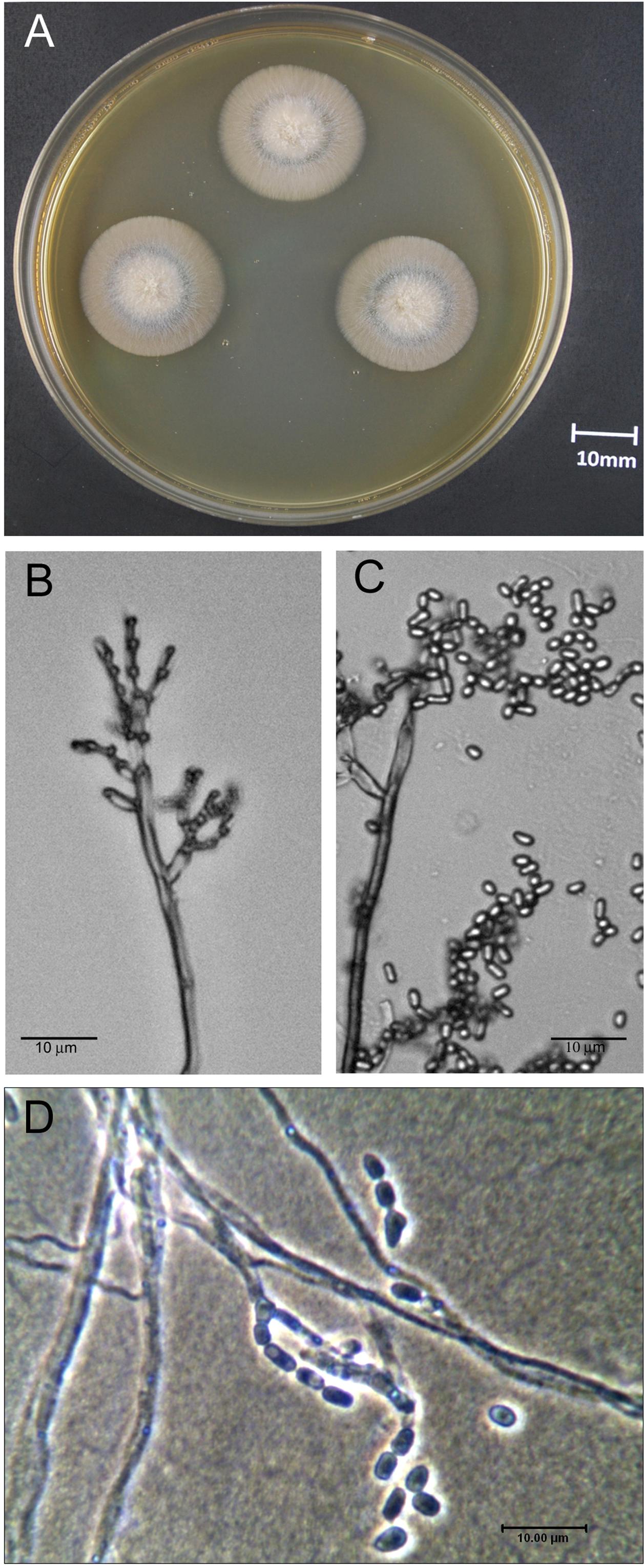
This technique provides the initial step in the process of analysing complex microbial communities in human, zoological and even geological settings. 
Broad-range 16S rDNA PCR is also more commonly used in research settings, originally for use in detecting and identifying unusual bacterial species but now more widely used in the rapidly expanding field of microbiome research. Although qPCR is by far the most frequently used molecular technique in bacterial diagnostics, in certain scenarios a broad-range (non-specific) 16S rDNA (ribosomal DNA) PCR is increasingly being used. At our hospital approximately 200 qPCRs are performed per week for the investigation of bacterial infections. The qPCR assay is a mainstay of microbiological diagnostics within the National Health Service (NHS). The technique uses specific primers (short strands of nucleic acid needed to initiate DNA replication) and fluorescent probes to allow real-time quantification of target bacterial DNA during amplification. 1 Quantitative PCR (qPCR), also known as specific PCR, involves the targeting of particular bacterial species. PCRs have revolutionised the detection of bacteria in clinical samples since their widespread introduction in the 1990s.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
